
Survival Analysis Step 3.0: Mediation Analysis
survival-analysis-step3.RmdWhat this step is for
This tutorial is the Step 3.0 survival-analysis article in
leo.ukb. After a main Cox model and any subgroup or
interaction work, a different question may arise:
does part of the exposure-outcome association operate through a mediator?
That is a mediation question, not a subgroup question and not an interaction question.
First separate mediation from interaction
The current survival-analysis workflow in leo.ukb
answers three different questions:
- Step 1: is there a main exposure-outcome association?
- Step 2: does that association differ across groups, and on which scale?
- Step 3.0: is part of that association transmitted through a mediator?
This distinction matters because:
- interaction is about effect heterogeneity
- mediation is about a pathway decomposition
In other words, mediation does not replace Step 1 or Step 2. It builds on them.
Start from the data and analytic roles
We use a synthetic example with a rare event, a binary exposure, a binary mediator, and several covariates.
set.seed(20260323)
n <- 500
age <- rnorm(n, 60, 8)
bmi <- rnorm(n, 26, 4)
smoking <- factor(sample(c("Never", "Ever"), n, replace = TRUE))
exposure_num <- rbinom(n, 1, plogis(-0.2 + 0.02 * (age - 60)))
mediator_num <- rbinom(n, 1, plogis(-0.5 + 0.9 * exposure_num + 0.02 * (bmi - 26)))
hazard_rate <- exp(
-5.8 + 0.55 * exposure_num + 0.40 * mediator_num +
0.02 * (age - 60) + 0.03 * (bmi - 26) + 0.20 * (smoking == "Ever")
)
event_time <- rexp(n, rate = hazard_rate)
censor_time <- rexp(n, rate = 0.08)
med_df <- data.frame(
outcome = as.integer(event_time <= censor_time),
outcome_censor = pmax(pmin(event_time, censor_time), 0.2),
exposure = factor(exposure_num, levels = c(0, 1), labels = c("Low risk", "High risk")),
mediator = factor(
mediator_num,
levels = c(0, 1),
labels = c("Low inflammation", "High inflammation")
),
age = age,
bmi = bmi,
smoking = smoking
)Before fitting anything, look at the analytic roles explicitly.
dplyr::glimpse(med_df)
#> Rows: 500
#> Columns: 7
#> $ outcome <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0…
#> $ outcome_censor <dbl> 2.6117515, 6.4186284, 8.6449270, 5.6413310, 12.1900483,…
#> $ exposure <fct> High risk, High risk, Low risk, High risk, Low risk, Lo…
#> $ mediator <fct> Low inflammation, High inflammation, Low inflammation, …
#> $ age <dbl> 59.73588, 59.19772, 45.83227, 57.49797, 46.83760, 55.76…
#> $ bmi <dbl> 27.17832, 27.91178, 30.48299, 24.70933, 22.63096, 28.43…
#> $ smoking <fct> Never, Never, Never, Never, Never, Ever, Ever, Ever, Ev…| Variable | Type | Step 3 role |
|---|---|---|
outcome |
Binary event indicator | Incident outcome event |
outcome_censor |
Follow-up time | Incident outcome follow-up time |
exposure |
Binary exposure | Exposure to be decomposed |
mediator |
Binary mediator | Candidate pathway variable |
age |
Continuous covariate | Adjustment covariate |
bmi |
Continuous covariate | Adjustment covariate |
smoking |
Binary covariate | Example covariate that requires explicit
x_cov_cond
|
Inspect the analysis-ready data before fitting
As in Step 1 and Step 2, it helps to inspect the analysis-ready cohort before moving to the model.
| outcome | outcome_censor | exposure | mediator | age | bmi | smoking |
|---|---|---|---|---|---|---|
| 0 | 2.611752 | High risk | Low inflammation | 59.73588 | 27.17832 | Never |
| 0 | 6.418628 | High risk | High inflammation | 59.19772 | 27.91178 | Never |
| 0 | 8.644927 | Low risk | Low inflammation | 45.83227 | 30.48299 | Never |
| 0 | 5.641331 | High risk | Low inflammation | 57.49797 | 24.70933 | Never |
| 0 | 12.190048 | Low risk | Low inflammation | 46.83760 | 22.63096 | Never |
| 0 | 20.568418 | Low risk | Low inflammation | 55.76899 | 28.43722 | Ever |
event_summary <- data.frame(
Metric = c("Participants", "Events", "Event rate"),
Value = c(
nrow(med_df),
sum(med_df$outcome),
sprintf("%.1f%%", 100 * mean(med_df$outcome))
)
)
knitr::kable(
event_summary,
caption = "Overall event burden in the mediation teaching data"
)| Metric | Value |
|---|---|
| Participants | 500 |
| Events | 30 |
| Event rate | 6.0% |
knitr::kable(
dplyr::count(med_df, exposure, mediator),
caption = "Exposure-mediator joint counts"
)| exposure | mediator | n |
|---|---|---|
| Low risk | Low inflammation | 163 |
| Low risk | High inflammation | 105 |
| High risk | Low inflammation | 102 |
| High risk | High inflammation | 130 |
These simple checks already answer three important questions:
- is the event rate low enough that
yreg = "auto"will reasonably choose the Cox path? - does the mediator distribution differ across exposure groups in a direction consistent with a possible pathway?
- do the covariates look interpretable enough to define a reproducible evaluation setting later?
Choose the mediation setup before fitting
Mediation analysis is easier to defend when the design choices are explicit before the model is run.
setup_table <- data.frame(
Component = c(
"Exposure contrast",
"Mediator type",
"Outcome model",
"Covariate evaluation profile"
),
Current_rule = c(
"Binary factor by default, or numeric with explicit x_exp_a0 and x_exp_a1",
"Binary mediator -> logistic; continuous/score-like mediator -> linear",
"yreg = 'auto' uses survCox when event rate <= 10%, otherwise survAFT_weibull",
"x_cov_cond should be explicit when x_cov contains non-numeric or binary numeric variables"
),
Why_it_matters = c(
"All direct and indirect effects are defined relative to this comparison",
"The mediator model determines how the pathway is represented",
"The final ratio scale depends on the actual outcome model",
"The decomposition is always evaluated at a specific covariate profile"
)
)
knitr::kable(
setup_table,
caption = "Choose the mediation setup before fitting"
)| Component | Current_rule | Why_it_matters |
|---|---|---|
| Exposure contrast | Binary factor by default, or numeric with explicit x_exp_a0 and x_exp_a1 | All direct and indirect effects are defined relative to this comparison |
| Mediator type | Binary mediator -> logistic; continuous/score-like mediator -> linear | The mediator model determines how the pathway is represented |
| Outcome model | yreg = ‘auto’ uses survCox when event rate <= 10%, otherwise survAFT_weibull | The final ratio scale depends on the actual outcome model |
| Covariate evaluation profile | x_cov_cond should be explicit when x_cov contains non-numeric or binary numeric variables | The decomposition is always evaluated at a specific covariate profile |
Know the method boundary before fitting
leo_cox_mediation() currently wraps
regmedint::regmedint() and keeps two model settings
parallel:
-
mregcontrols the mediator model, i.e.x_med ~ x_exp + x_cov -
yregcontrols the outcome model, i.e.time ~ x_exp + x_med + x_cov - when
interaction = TRUE, the outcome model becomestime ~ x_exp + x_med + x_exp:x_med + x_cov
The current wrapper is most appropriate when:
- the outcome is time-to-event
- the exposure is binary, or numeric with explicit
x_exp_a0andx_exp_a1 - the mediator is binary or continuous
- the temporal ordering is scientifically defensible
Two boundaries are especially important in UK Biobank-style analyses that rely heavily on baseline measurements:
- if the final
yreg(the outcome-model setting here) issurvCox, the decomposition relies on the rare-event approximation; in plain terms, this means the split is easier to defend as an approximate result when events are relatively rare, rather than as an exact estimate - if exposure and mediator are measured at the same baseline visit, the data usually cannot show that exposure came before mediator, so the result is better written as decomposition evidence that is consistent with an assumed pathway; without external evidence on temporal ordering, it should not be written up as a strong causal claim
Fit the simplest mediation workflow
As in the Cox tutorials, define the adjustment hierarchy first.
med_models <- list(
"Model A" = c("age"),
"Model B" = c("age", "bmi")
)
res_med <- leo_cox_mediation(
df = med_df,
y_out = c("outcome", "outcome_censor"),
x_exp = "exposure",
x_med = "mediator",
x_cov = med_models,
verbose = FALSE
)
res_med$result
#> Model Effect code Effect Scale Exposure
#> 1 Model A cde Controlled direct effect Hazard ratio exposure
#> 2 Model A pnde Pure natural direct effect Hazard ratio exposure
#> 3 Model A tnie Total natural indirect effect Hazard ratio exposure
#> 4 Model A tnde Total natural direct effect Hazard ratio exposure
#> 5 Model A pnie Pure natural indirect effect Hazard ratio exposure
#> 6 Model A te Total effect Hazard ratio exposure
#> 7 Model A pm Proportion mediated Proportion exposure
#> 8 Model B cde Controlled direct effect Hazard ratio exposure
#> 9 Model B pnde Pure natural direct effect Hazard ratio exposure
#> 10 Model B tnie Total natural indirect effect Hazard ratio exposure
#> 11 Model B tnde Total natural direct effect Hazard ratio exposure
#> 12 Model B pnie Pure natural indirect effect Hazard ratio exposure
#> 13 Model B te Total effect Hazard ratio exposure
#> 14 Model B pm Proportion mediated Proportion exposure
#> Mediator Outcome Exposure contrast Mediator reference N Case N
#> 1 mediator outcome Low risk -> High risk High inflammation 500 30
#> 2 mediator outcome Low risk -> High risk <NA> 500 30
#> 3 mediator outcome Low risk -> High risk <NA> 500 30
#> 4 mediator outcome Low risk -> High risk <NA> 500 30
#> 5 mediator outcome Low risk -> High risk <NA> 500 30
#> 6 mediator outcome Low risk -> High risk <NA> 500 30
#> 7 mediator outcome Low risk -> High risk <NA> 500 30
#> 8 mediator outcome Low risk -> High risk High inflammation 500 30
#> 9 mediator outcome Low risk -> High risk <NA> 500 30
#> 10 mediator outcome Low risk -> High risk <NA> 500 30
#> 11 mediator outcome Low risk -> High risk <NA> 500 30
#> 12 mediator outcome Low risk -> High risk <NA> 500 30
#> 13 mediator outcome Low risk -> High risk <NA> 500 30
#> 14 mediator outcome Low risk -> High risk <NA> 500 30
#> Non-event N Total follow-up Estimate 95% CI P value Mediator model
#> 1 470 5470.759 1.872 0.639, 5.479 0.253 logistic
#> 2 470 5470.759 1.443 0.690, 3.018 0.329 logistic
#> 3 470 5470.759 1.058 0.891, 1.258 0.518 logistic
#> 4 470 5470.759 1.557 0.738, 3.281 0.245 logistic
#> 5 470 5470.759 0.981 0.817, 1.179 0.842 logistic
#> 6 470 5470.759 1.528 0.743, 3.140 0.249 logistic
#> 7 470 5470.759 0.160 -0.356, 0.676 0.543 logistic
#> 8 470 5470.759 1.938 0.661, 5.678 0.228 logistic
#> 9 470 5470.759 1.470 0.698, 3.096 0.311 logistic
#> 10 470 5470.759 1.054 0.870, 1.276 0.593 logistic
#> 11 470 5470.759 1.600 0.761, 3.366 0.215 logistic
#> 12 470 5470.759 0.968 0.791, 1.185 0.753 logistic
#> 13 470 5470.759 1.549 0.753, 3.185 0.234 logistic
#> 14 470 5470.759 0.144 -0.402, 0.689 0.606 logistic
#> Outcome model
#> 1 survCox
#> 2 survCox
#> 3 survCox
#> 4 survCox
#> 5 survCox
#> 6 survCox
#> 7 survCox
#> 8 survCox
#> 9 survCox
#> 10 survCox
#> 11 survCox
#> 12 survCox
#> 13 survCox
#> 14 survCoxHow to read the result table
The mediation output is easier to read if you separate the columns into roles:
-
Effect codeandEffectidentify which decomposition quantity is being shown -
Scalereminds you thatpmis a proportion, whereas the other rows are ratio measures -
Exposure contrastshows thex_exp_a0 -> x_exp_a1comparison -
Mediator referenceis populated for thecderow and echoed in$evaluation -
Mediator modelandOutcome modelshow the actualmregandyregselected
The scientific point is simple:
mediation estimates are not free-floating numbers; they are conditional on a chosen exposure contrast, mediator reference, covariate profile, and model pair
Teach the core settings in leo.ukb language
regmedint uses a compact set of evaluation settings and
model settings. leo_cox_mediation() keeps the same
scientific meaning while presenting them in more readable
leo.ukb language.
| Question |
leo.ukb parameter |
What it means in practice |
|---|---|---|
| Which exposure contrast is being compared? |
x_exp_a0, x_exp_a1
|
The exposure comparison used to define the mediation effect, such as
Low risk -> High risk or
8.0 -> 9.0. |
| At what mediator value is the controlled direct effect evaluated? | x_med_cde |
The mediator reference value used for cde. |
| Under what covariate profile are the effects evaluated? | x_cov_cond |
The covariate setting at which the effects are computed. |
| How is the mediator modeled? | mreg |
The mediator model for x_med ~ x_exp + x_cov. |
| How is the survival outcome modeled? | yreg |
The outcome model for
time ~ x_exp + x_med + x_cov. |
Should the outcome model include x_exp:x_med? |
interaction |
If TRUE, the outcome model allows the effect of
x_exp to vary with x_med. |
In other words:
-
x_exp_a0,x_exp_a1,x_med_cde, andx_cov_conddefine where the mediation effects are evaluated -
mreg,yreg, andinteractiondefine which models generate that decomposition
How mreg = "auto" behaves
The current leo.ukb rule is:
- binary factor / character / logical mediator ->
logistic - two-valued integer-like numeric mediator such as
0/1->logistic - numeric mediator with more than two observed values, or non-integer
two-valued coding such as
0.2/0.8->linear - multi-level categorical mediator stored as factor -> not currently supported
This is the practical interpretation:
- use
autofreely for binary mediators - use
autofor true continuous biomarkers - for ordered scores, such as a low-grade inflammation index from
-7to7, keep the variable numeric and usually let it behave as a continuous mediator - if the scientific question is threshold-based, dichotomize first and then use a logistic mediator model
Decode the abbreviations before interpreting anything
data.frame(
`Effect code` = c("cde", "pnde", "tnie", "tnde", "pnie", "te", "pm"),
Meaning = c(
"Controlled direct effect",
"Pure natural direct effect",
"Total natural indirect effect",
"Total natural direct effect",
"Pure natural indirect effect",
"Total effect",
"Proportion mediated"
),
`Practical reading` = c(
"Direct effect when the mediator is fixed at the chosen reference value.",
"Direct pathway not operating through the mediator.",
"Indirect pathway operating through the mediator.",
"Direct pathway allowing the mediator to vary naturally.",
"Indirect pathway under the pure natural definition.",
"Overall exposure-outcome effect.",
"Share of the total effect attributed to the mediator pathway."
)
)
#> Effect.code Meaning
#> 1 cde Controlled direct effect
#> 2 pnde Pure natural direct effect
#> 3 tnie Total natural indirect effect
#> 4 tnde Total natural direct effect
#> 5 pnie Pure natural indirect effect
#> 6 te Total effect
#> 7 pm Proportion mediated
#> Practical.reading
#> 1 Direct effect when the mediator is fixed at the chosen reference value.
#> 2 Direct pathway not operating through the mediator.
#> 3 Indirect pathway operating through the mediator.
#> 4 Direct pathway allowing the mediator to vary naturally.
#> 5 Indirect pathway under the pure natural definition.
#> 6 Overall exposure-outcome effect.
#> 7 Share of the total effect attributed to the mediator pathway.For most applied readers, three rows are especially important:
-
te: how large the overall association is -
tnie: how large the mediator pathway appears to be -
pm: what fraction of the total effect may be operating through the mediator
Turn the result into a clinically readable summary
key_rows <- dplyr::filter(
res_med$result,
Model == "Model B",
`Effect code` %in% c("te", "tnie", "pm")
)
key_rows
#> Model Effect code Effect Scale Exposure
#> 1 Model B tnie Total natural indirect effect Hazard ratio exposure
#> 2 Model B te Total effect Hazard ratio exposure
#> 3 Model B pm Proportion mediated Proportion exposure
#> Mediator Outcome Exposure contrast Mediator reference N Case N
#> 1 mediator outcome Low risk -> High risk <NA> 500 30
#> 2 mediator outcome Low risk -> High risk <NA> 500 30
#> 3 mediator outcome Low risk -> High risk <NA> 500 30
#> Non-event N Total follow-up Estimate 95% CI P value Mediator model
#> 1 470 5470.759 1.054 0.870, 1.276 0.593 logistic
#> 2 470 5470.759 1.549 0.753, 3.185 0.234 logistic
#> 3 470 5470.759 0.144 -0.402, 0.689 0.606 logistic
#> Outcome model
#> 1 survCox
#> 2 survCox
#> 3 survCox
te_row <- dplyr::filter(key_rows, `Effect code` == "te")
tnie_row <- dplyr::filter(key_rows, `Effect code` == "tnie")
pm_row <- dplyr::filter(key_rows, `Effect code` == "pm")
pm_ci <- as.numeric(trimws(strsplit(pm_row$`95% CI`, ",", fixed = TRUE)[[1]]))
ratio_label <- if (te_row$`Outcome model` == "survCox") "HR" else "TR"
ratio_noun <- if (te_row$`Outcome model` == "survCox") "hazard" else "time"
te_pct <- 100 * (te_row$Estimate - 1)
pm_pct <- 100 * pm_row$Estimate
data.frame(
Quantity = c("Total effect", "Indirect effect through inflammation", "Proportion mediated"),
`Model B summary` = c(
sprintf("%s %.3f (%s)", ratio_label, te_row$Estimate, te_row$`95% CI`),
sprintf("%s %.3f (%s)", ratio_label, tnie_row$Estimate, tnie_row$`95% CI`),
sprintf("%.1f%% (%.1f%% to %.1f%%)", 100 * pm_row$Estimate, 100 * pm_ci[1], 100 * pm_ci[2])
)
)
#> Quantity Model.B.summary
#> 1 Total effect HR 1.549 (0.753, 3.185)
#> 2 Indirect effect through inflammation HR 1.054 (0.870, 1.276)
#> 3 Proportion mediated 14.4% (-40.2% to 68.9%)Because the event rate in this example is only 6.0%,
yreg = "auto" selected survCox, so the ratio
scale is a hazard ratio.
In manuscript-style language, the Model B result can be read as:
- the
High riskgroup has an overall hazard ratio that is about 54.9% higher than theLow riskgroup (te = 1.549) - the inflammation pathway contributes only a modest indirect effect
(
tnie = 1.054) - the point estimate for the proportion mediated is 14.4%, suggesting that part of the overall association could be operating through the inflammation pathway
The critical scientific qualifier is uncertainty. Here the confidence
interval for pm is wide and crosses zero, so the careful
interpretation is not “mediation has been established”, but:
the estimates are compatible with partial mediation through inflammation, but the pathway effect remains too imprecise to support a strong mechanistic claim on its own
Inspect the evaluation settings directly
leo_cox_mediation() keeps a compact evaluation summary
in $evaluation.
res_med$evaluation
#> model exposure mediator outcome n case_n non_event_n followup_total
#> 1 Model A exposure mediator outcome 500 30 470 5470.759
#> 2 Model B exposure mediator outcome 500 30 470 5470.759
#> event_rate exposure_contrast exposure_contrast_value x_exp_a0 x_exp_a1
#> 1 0.06 Low risk -> High risk 0.000 -> 1.000 0 1
#> 2 0.06 Low risk -> High risk 0.000 -> 1.000 0 1
#> mediator_reference x_med_cde x_med_cde_internal mreg yreg
#> 1 High inflammation High inflammation 1 logistic survCox
#> 2 High inflammation High inflammation 1 logistic survCox
#> pm_unstable
#> 1 TRUE
#> 2 TRUE
#> pm_note
#> 1 Total effect 95% CI includes the null value; proportion mediated (pm) may be unstable and should be interpreted cautiously.
#> 2 Total effect 95% CI includes the null value; proportion mediated (pm) may be unstable and should be interpreted cautiously.
#> interaction covariates x_cov_cond
#> 1 TRUE age age=59.523
#> 2 TRUE age, bmi age=59.523; bmi=25.464This table is especially useful when you want to audit:
- which
x_exp_a0 -> x_exp_a1contrast was used - which mediator reference was used for the controlled direct effect
- which covariate profile was used for
x_cov_cond - which
mregandyregwere used - whether the outcome model included
x_exp:x_med
Non-numeric covariates need explicit x_cov_cond
When the selected adjustment set contains a non-numeric covariate,
the evaluation profile must be given explicitly. The same applies to
binary numeric covariates such as 0/1 indicators, because
the default median can otherwise create impossible values such as
0.5.
res_med_smoking <- leo_cox_mediation(
df = med_df,
y_out = c("outcome", "outcome_censor"),
x_exp = "exposure",
x_med = "mediator",
x_cov = "smoking",
x_cov_cond = list(smoking = "Never"),
verbose = FALSE
)
res_med_smoking$evaluation
#> model exposure mediator outcome n case_n non_event_n followup_total
#> 1 model_1 exposure mediator outcome 500 30 470 5470.759
#> event_rate exposure_contrast exposure_contrast_value x_exp_a0 x_exp_a1
#> 1 0.06 Low risk -> High risk 0.000 -> 1.000 0 1
#> mediator_reference x_med_cde x_med_cde_internal mreg yreg
#> 1 High inflammation High inflammation 1 logistic survCox
#> pm_unstable
#> 1 TRUE
#> pm_note
#> 1 Total effect 95% CI includes the null value; proportion mediated (pm) may be unstable and should be interpreted cautiously.
#> interaction covariates x_cov_cond
#> 1 TRUE smoking smoking=NeverThis is not just more transparent. It is required whenever
x_cov contains a factor or binary numeric covariate,
because the evaluation profile has to map to a real person that the
reader can reconstruct.
Continuous mediators, including score-like mediators, are also supported
The same workflow can also be used with a continuous mediator.
med_df_cont <- dplyr::mutate(
med_df,
mediator_cont = rnorm(
n,
mean = 0.5 * as.integer(exposure == "High risk") + 0.02 * (age - 60),
sd = 1
)
)
res_med_cont <- leo_cox_mediation(
df = med_df_cont,
y_out = c("outcome", "outcome_censor"),
x_exp = "exposure",
x_med = "mediator_cont",
x_cov = "age",
mreg = "linear",
verbose = FALSE
)
res_med_cont$result
#> Model Effect code Effect Scale Exposure
#> 1 model_1 cde Controlled direct effect Hazard ratio exposure
#> 2 model_1 pnde Pure natural direct effect Hazard ratio exposure
#> 3 model_1 tnie Total natural indirect effect Hazard ratio exposure
#> 4 model_1 tnde Total natural direct effect Hazard ratio exposure
#> 5 model_1 pnie Pure natural indirect effect Hazard ratio exposure
#> 6 model_1 te Total effect Hazard ratio exposure
#> 7 model_1 pm Proportion mediated Proportion exposure
#> Mediator Outcome Exposure contrast Mediator reference N Case N
#> 1 mediator_cont outcome Low risk -> High risk 0.233 500 30
#> 2 mediator_cont outcome Low risk -> High risk <NA> 500 30
#> 3 mediator_cont outcome Low risk -> High risk <NA> 500 30
#> 4 mediator_cont outcome Low risk -> High risk <NA> 500 30
#> 5 mediator_cont outcome Low risk -> High risk <NA> 500 30
#> 6 mediator_cont outcome Low risk -> High risk <NA> 500 30
#> 7 mediator_cont outcome Low risk -> High risk <NA> 500 30
#> Non-event N Total follow-up Estimate 95% CI P value Mediator model
#> 1 470 5470.759 1.562 0.760, 3.213 0.225 linear
#> 2 470 5470.759 1.694 0.801, 3.584 0.168 linear
#> 3 470 5470.759 0.917 0.775, 1.086 0.315 linear
#> 4 470 5470.759 1.554 0.747, 3.233 0.238 linear
#> 5 470 5470.759 1.000 0.849, 1.178 0.999 linear
#> 6 470 5470.759 1.554 0.755, 3.201 0.231 linear
#> 7 470 5470.759 -0.252 -0.849, 0.345 0.407 linear
#> Outcome model
#> 1 survCox
#> 2 survCox
#> 3 survCox
#> 4 survCox
#> 5 survCox
#> 6 survCox
#> 7 survCoxFor a continuous mediator, the cde row shows a numeric
mediator reference because the controlled direct effect is evaluated at
x_med_cde.
This is also the workflow we would usually recommend for an ordered
score-like mediator. For example, if a low-grade inflammation index
ranges from -7 to 7, the first question is
scientific rather than computational:
- does the index behave like a severity score, where one-unit changes are interpretable enough for a linear approximation?
- or is the real question about a thresholded state that should be defined as binary first?
For most ordered severity scores, the cleaner first pass is to keep the score numeric and fit a linear mediator model.
Add a paper-style pathway figure
For a single mediator, a compact pathway figure often communicates the result faster than a dense table alone.
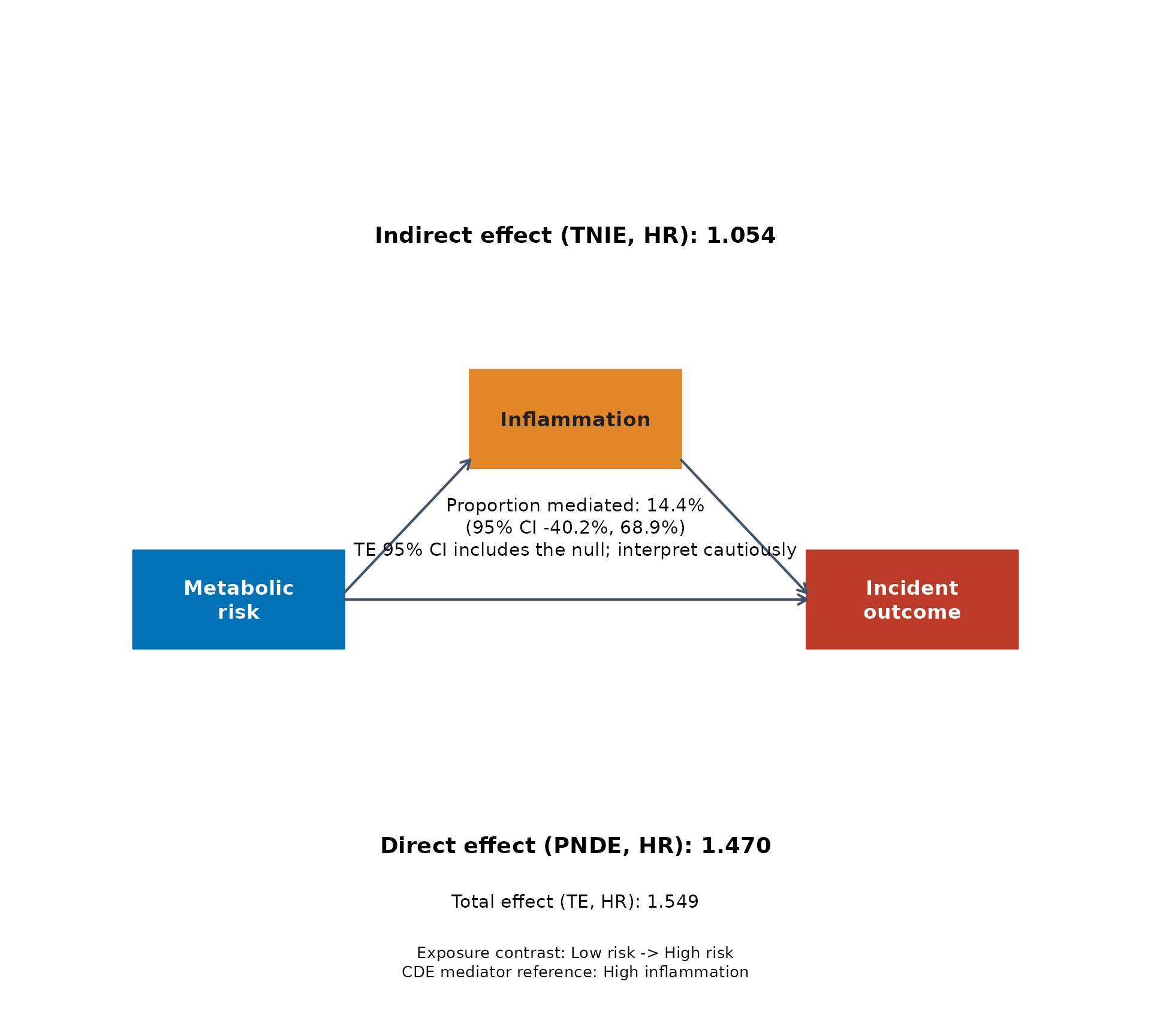
leo_cox_mediation_plot(
x = res_med,
model = "Model B",
exposure_label = "Metabolic\nrisk",
mediator_label = "Inflammation",
outcome_label = "Incident\noutcome",
language = "en",
palette = "jama"
)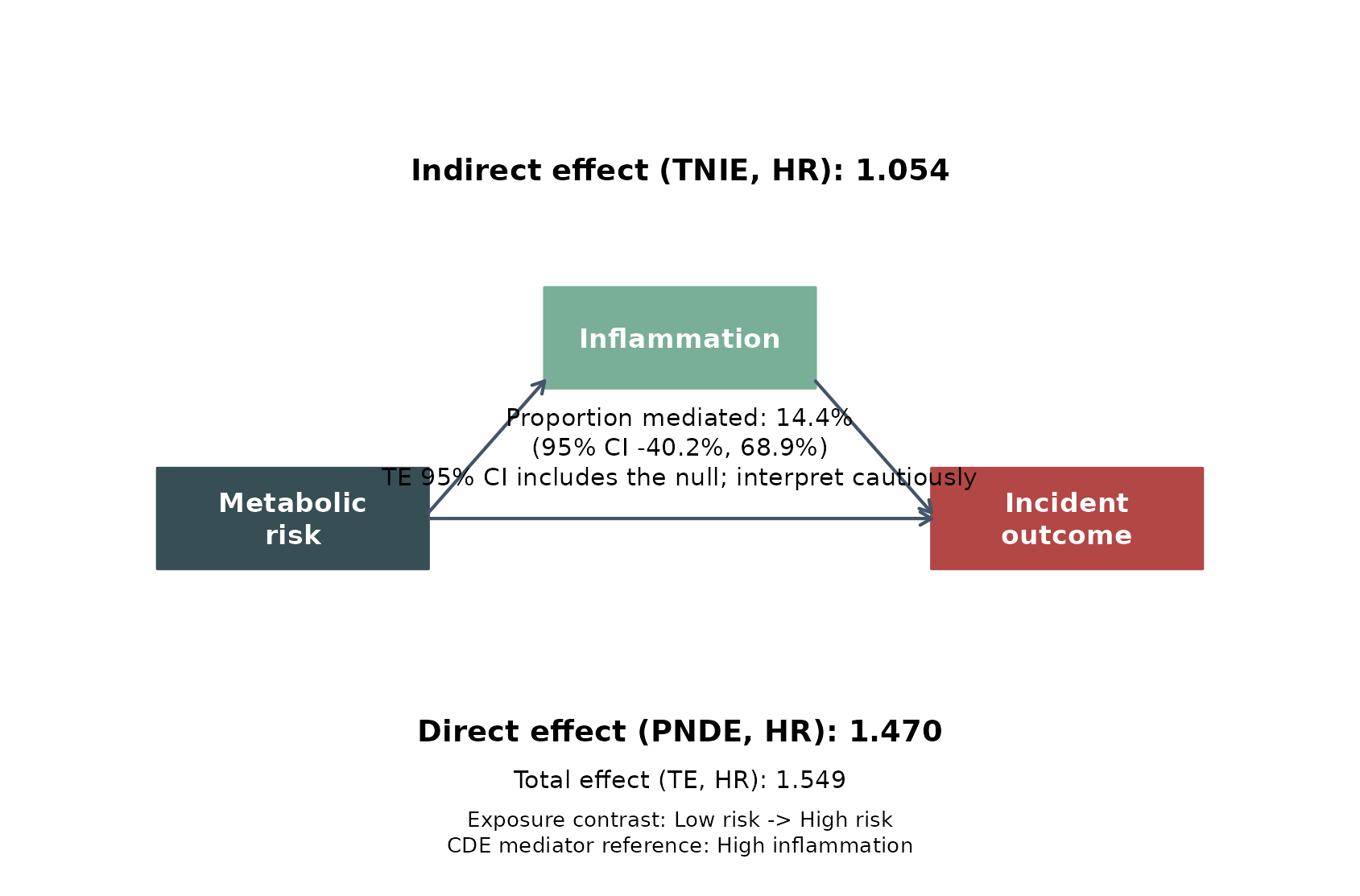
This figure is useful because it keeps the main teaching elements in one place:
- left box: the exposure contrast
- middle box: the proposed mediator
- right box: the incident outcome
- top text: the indirect pathway through the mediator
- middle text: the estimated proportion of the total effect attributed to that pathway
- bottom text: the direct pathway not explained by the mediator
One subtle but important point is that the mediated pathway shown
here is the full X -> M -> Y pathway. The figure does
not print a separate M -> Y coefficient because
regmedint reports causal decomposition quantities such as
tnie, pnde, and te, rather than a
simple product of two regression coefficients.
Compare the built-in journal palettes
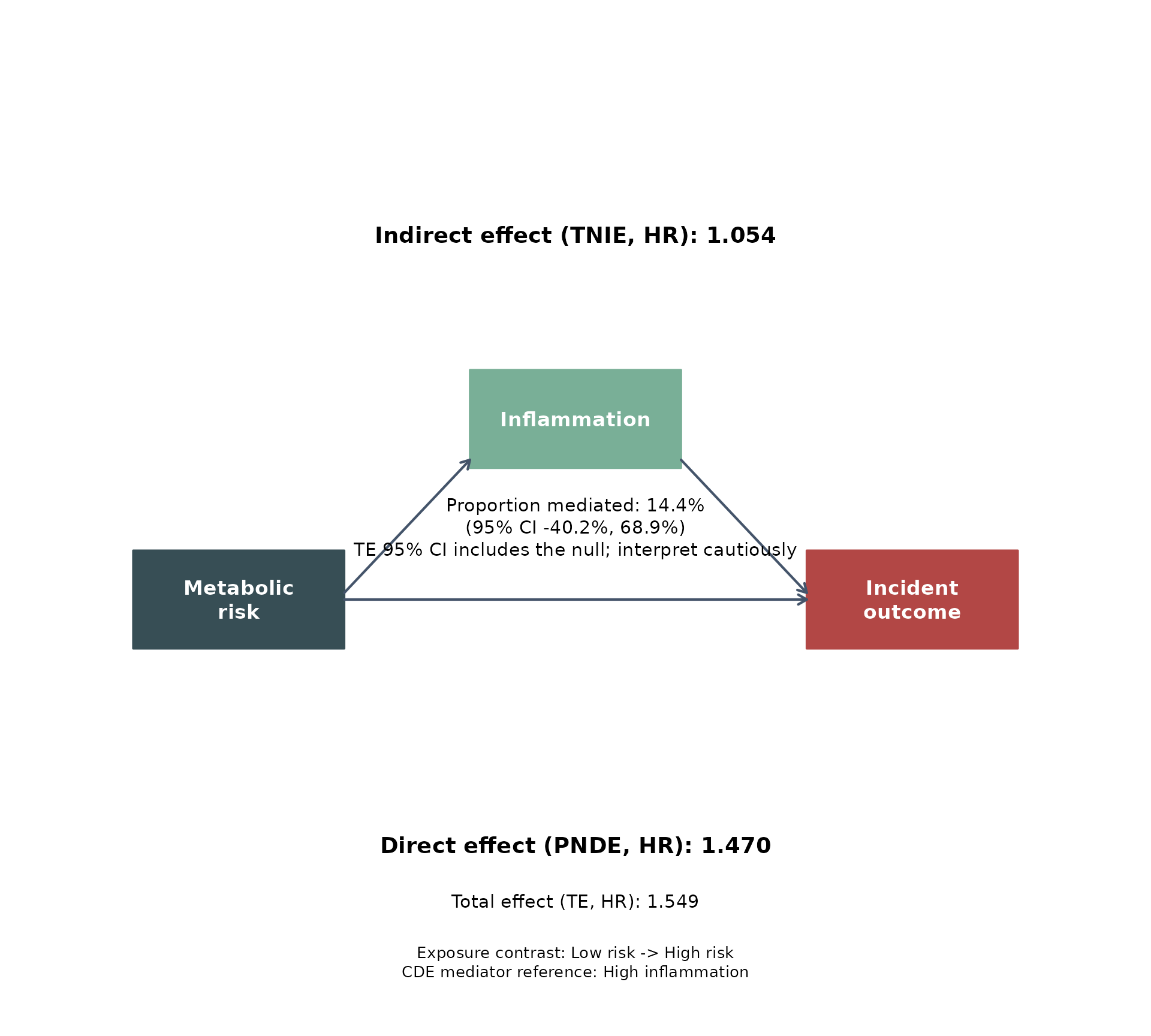
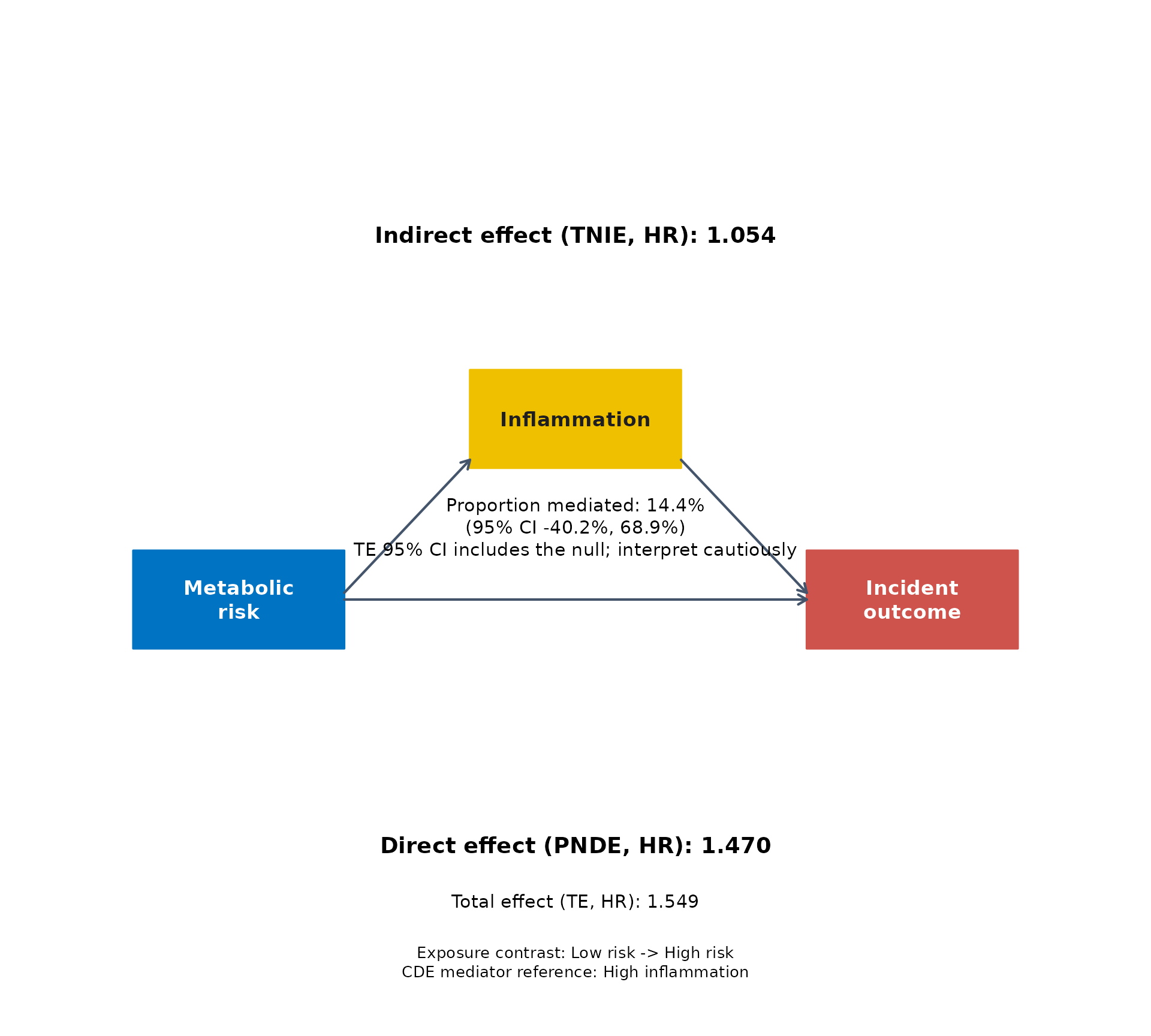
The same fitted result can be displayed with several built-in journal-style palettes.
for (pal in c("jama", "jco", "lancet", "nejm")) {
leo_cox_mediation_plot(
x = res_med,
model = "Model B",
exposure_label = "Metabolic\nrisk",
mediator_label = "Inflammation",
outcome_label = "Incident\noutcome",
language = "en",
palette = pal
)
}
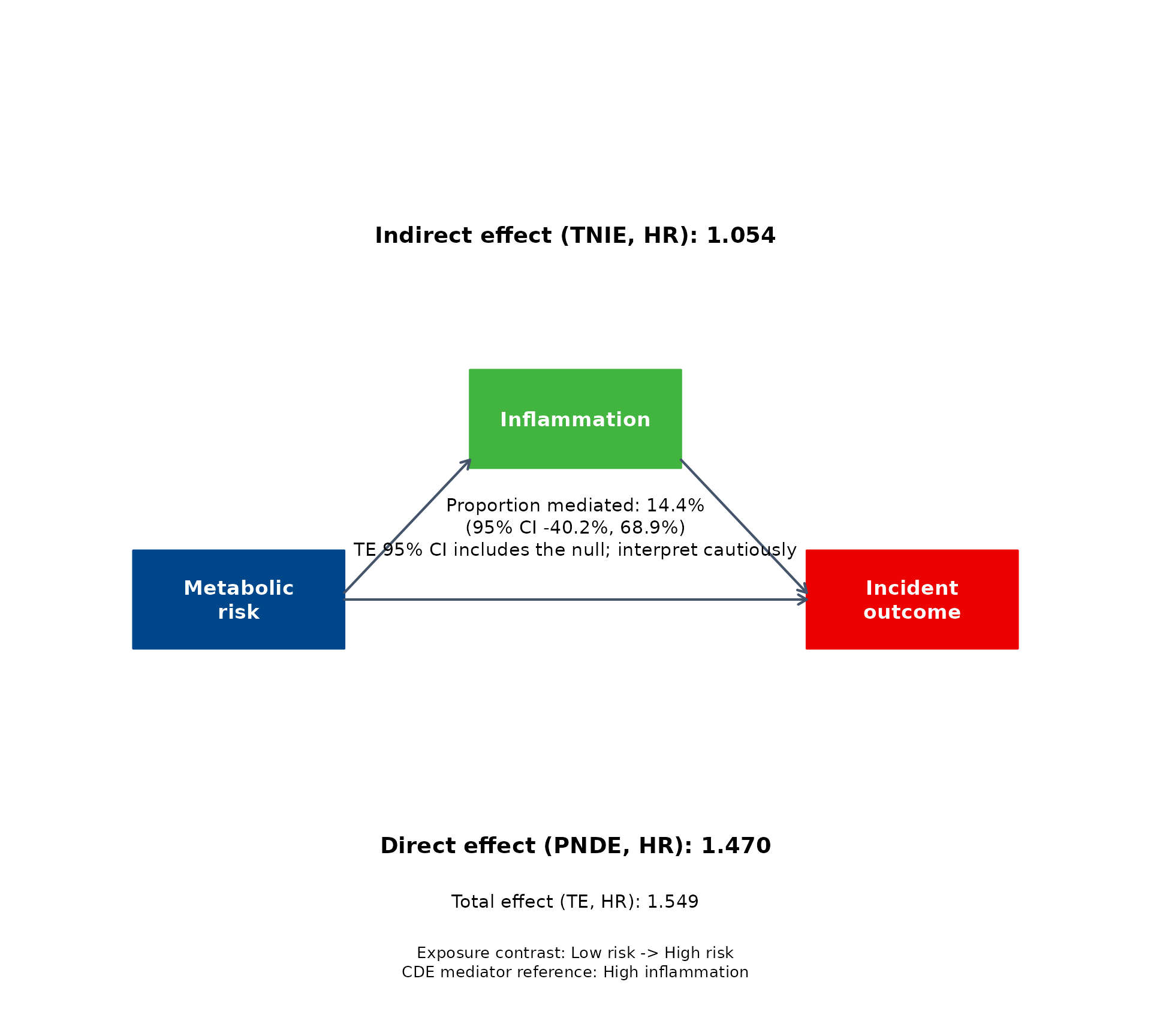
How to interpret common result patterns
In applied survival mediation analysis, the most common interpretation patterns are:
-
teis clearly away from the null,tnieis modest, andpmis positive but imprecise: compatible with partial mediation, but not decisive -
teandtnieare both clearly away from the null, andpmis precise: stronger evidence that the pathway may explain a meaningful share of the total effect -
teis weak or crosses the null:pmcan become unstable and should be treated cautiously -
yregswitches to AFT: all non-pmeffect rows should now be interpreted as time ratios rather than hazard ratios
This is why Step 3 is not only about producing a decomposition table. It is also about learning when the estimates are scientifically interpretable and when they are still too unstable for a strong claim.
How to write this in a paper
A defensible reporting order is usually:
- report the main exposure-outcome association first
- then present the mediation analysis as a pathway decomposition
- state the exposure contrast, mediator reference, and adjustment profile
- report which
yregwas used; if it wassurvCox, mention the rare-event approximation and make clear that this means the decomposition is only being reported as an approximate split when events are relatively rare, and if it was AFT, remember that the ratio scale is a time ratio - if exposure and mediator are measured at the same baseline visit, write the result up as decomposition evidence that is consistent with an assumed pathway, and say explicitly that the data alone do not establish exposure before mediator unless temporal ordering is supported externally
In other words, mediation should usually refine the biological story after the main Cox result is already established.
What to take away from Step 3
This step should leave you with six practical rules:
- mediation asks a pathway question, not a heterogeneity question
- the data should be checked before fitting, just as in Step 1 and Step 2
-
x_exp_a0,x_exp_a1,x_med_cde, andx_cov_conddefine where the mediation effects are evaluated -
mregcontrolsx_med ~ x_exp + x_cov,yregcontrolstime ~ x_exp + x_med + x_cov, andinteraction = TRUEaddsx_exp:x_medto the outcome model -
yreg = "auto"choosessurvCoxfor event proportion<= 10%(which means the decomposition then relies on the rare-event approximation and is better read as approximate when events are relatively rare) andsurvAFT_weibullotherwise - factor mediators should not be forced into
mreg = "linear", and the final mediation result should always be interpreted with the actual outcome model, uncertainty, and the study’s temporal ordering in mind