Cox mediation analysis
leo_cox_mediation.Rdleo_cox_mediation_plot() turns a leo_cox_mediation result into a compact
pathway figure for a selected model. The plot keeps all annotation text on a
shared vertical center axis, uses equal-sized boxes, and draws only straight
arrows between the exposure, mediator, and outcome nodes.
Usage
leo_cox_mediation(
df,
y_out,
x_exp,
x_med,
x_cov = NULL,
event_value = 1,
min_followup_time = 0,
x_exp_a0 = NULL,
x_exp_a1 = NULL,
x_med_cde = NULL,
x_cov_cond = NULL,
mreg = c("auto", "linear", "logistic"),
yreg = c("auto", "survCox", "survAFT_weibull", "survAFT_exp"),
interaction = TRUE,
verbose = TRUE
)
leo_cox_mediation_plot(
x,
model = NULL,
exposure_label = "Exposure",
mediator_label = "Mediator",
outcome_label = "Outcome",
language = c("en", "zh"),
palette = c("jama", "jco", "lancet", "nejm"),
add_note = TRUE,
font_family = NULL
)Arguments
- df
Data frame with outcome, time, exposure, mediator, and covariates.
- y_out
c(event, time)column names.- x_exp
Exposure column name.
- x_med
Mediator column name.
- x_cov
Covariate column names, or
NULL.- event_value
Value indicating an event.
- min_followup_time
Minimum follow-up time filter.
- x_exp_a0
Reference value of
x_exp. Defaults to first factor level; required if x_exp is character (not factor). For a binary factor exposure, either use the default contrast or supplyx_exp_a0andx_exp_a1together. For numericx_exp, if exactly two unique observed values remain after filtering, they are used as the default contrast; otherwise, supplyx_exp_a0andx_exp_a1explicitly.- x_exp_a1
Contrasted value of
x_exp. Defaults to second factor level; required if x_exp is character (not factor). For a binary factor exposure, either use the default contrast or supplyx_exp_a0andx_exp_a1together. For numericx_exp, if exactly two unique observed values remain after filtering, they are used as the default contrast; otherwise, supplyx_exp_a0andx_exp_a1explicitly.- x_med_cde
Value of
x_medfor CDE evaluation. Defaults to second factor level (binary) or median (continuous); required if x_med is character (not factor). For logistic mediators, use the original observed mediator value rather than an internal 0/1 recode unless the observed values are themselves 0/1.- x_cov_cond
Covariate values for effect evaluation. Named list/vector for non-numeric covariates; defaults to medians. Required for factor/binary covariates.
- mreg
Mediator model:
"auto","linear", or"logistic".- yreg
Outcome model:
"auto","survCox","survAFT_weibull", or"survAFT_exp".- interaction
Include
x_exp:x_medinteraction? IfTRUE, the outcome model addsx_exp:x_med, i.e.time ~ x_exp + x_med + x_exp:x_med + x_cov.- verbose
Print progress messages.
- x
A
leo_cox_mediationobject returned byleo_cox_mediation().- model
Character scalar giving the model to display. Defaults to the last model in
x$result.- exposure_label
Character label shown in the left box. Manual
\nline breaks are respected.- mediator_label
Character label shown in the middle box. Manual
\nline breaks are respected.- outcome_label
Character label shown in the right box. Manual
\nline breaks are respected.- language
One of
"en"or"zh".- palette
One of
"jama","jco","lancet", or"nejm".- add_note
Logical; whether to show tutorial-style note text such as the exposure contrast, CDE mediator reference, and a TE-null caution when applicable.
- font_family
Optional graphics font family. Use generic families such as
"sans","serif", or"mono"for the most robust cross-device output.
Value
leo_cox_mediation object with $result, $result_detail, $evaluation, $fit.
For leo_cox_mediation(), a leo_cox_mediation object containing a display table in $result, a detailed mediation table in $result_detail, an evaluation summary in $evaluation, and fitted regmedint objects in $fit. For leo_cox_mediation_plot(), an invisible recordedplot object after drawing the pathway figure on the active graphics device.
Details
Performs causal mediation analysis for survival outcomes via regmedint::regmedint().
Fits mediator model x_med ~ x_exp + x_cov and outcome model
time ~ x_exp + x_med + x_cov (with optional x_exp:x_med interaction).
yreg = "auto" uses the observed event proportion in the post-filter,
complete-case analysis set: Cox for rare outcomes (<=10% events), AFT otherwise.
This 10% threshold is an empirical rare-event approximation rule.
Examples
if (requireNamespace("regmedint", quietly = TRUE)) {
set.seed(123)
n <- 200
age <- rnorm(n, 60, 8)
exposure <- rbinom(n, 1, 0.5)
mediator <- rnorm(n, 0.6 * exposure + 0.02 * (age - 60), 1)
time_event <- rexp(n, rate = exp(-5.8 + 0.55 * exposure + 0.20 * mediator + 0.02 * (age - 60)))
time_censor <- rexp(n, rate = 0.08)
med_df <- data.frame(
outcome = as.integer(time_event <= time_censor),
outcome_censor = pmax(pmin(time_event, time_censor), 0.1),
exposure = exposure, mediator = mediator, age = age
)
res_med <- leo_cox_mediation(
df = med_df, y_out = c("outcome", "outcome_censor"),
x_exp = "exposure", x_med = "mediator", x_cov = "age",
verbose = FALSE
)
res_med$result
}
#> Model Effect code Effect Scale Exposure
#> 1 model_1 cde Controlled direct effect Hazard ratio exposure
#> 2 model_1 pnde Pure natural direct effect Hazard ratio exposure
#> 3 model_1 tnie Total natural indirect effect Hazard ratio exposure
#> 4 model_1 tnde Total natural direct effect Hazard ratio exposure
#> 5 model_1 pnie Pure natural indirect effect Hazard ratio exposure
#> 6 model_1 te Total effect Hazard ratio exposure
#> 7 model_1 pm Proportion mediated Proportion exposure
#> Mediator Outcome Exposure contrast Mediator reference N Case N Non-event N
#> 1 mediator outcome 0.000 -> 1.000 0.370 200 8 192
#> 2 mediator outcome 0.000 -> 1.000 <NA> 200 8 192
#> 3 mediator outcome 0.000 -> 1.000 <NA> 200 8 192
#> 4 mediator outcome 0.000 -> 1.000 <NA> 200 8 192
#> 5 mediator outcome 0.000 -> 1.000 <NA> 200 8 192
#> 6 mediator outcome 0.000 -> 1.000 <NA> 200 8 192
#> 7 mediator outcome 0.000 -> 1.000 <NA> 200 8 192
#> Total follow-up Estimate 95% CI P value Mediator model Outcome model
#> 1 2545.621 3.247 0.540, 19.534 0.198 linear survCox
#> 2 2545.621 3.420 0.626, 18.687 0.156 linear survCox
#> 3 2545.621 1.376 0.865, 2.189 0.178 linear survCox
#> 4 2545.621 3.741 0.570, 24.566 0.169 linear survCox
#> 5 2545.621 1.258 0.567, 2.788 0.572 linear survCox
#> 6 2545.621 4.705 0.865, 25.595 0.073 linear survCox
#> 7 2545.621 0.347 -0.091, 0.784 0.120 linear survCox
# Continuous exposure requires x_exp_a0 and x_exp_a1
if (requireNamespace("regmedint", quietly = TRUE)) {
set.seed(123)
n <- 200
cont_df <- data.frame(
outcome = rbinom(n, 1, 0.06),
outcome_censor = rexp(n, 0.1),
exposure = rnorm(n), mediator = rnorm(n), age = rnorm(n, 60, 8)
)
res_cont <- leo_cox_mediation(
df = cont_df, y_out = c("outcome", "outcome_censor"),
x_exp = "exposure", x_med = "mediator", x_cov = "age",
x_exp_a0 = -1, x_exp_a1 = 1, verbose = FALSE
)
res_cont$result
}
#> Model Effect code Effect Scale Exposure
#> 1 model_1 cde Controlled direct effect Hazard ratio exposure
#> 2 model_1 pnde Pure natural direct effect Hazard ratio exposure
#> 3 model_1 tnie Total natural indirect effect Hazard ratio exposure
#> 4 model_1 tnde Total natural direct effect Hazard ratio exposure
#> 5 model_1 pnie Pure natural indirect effect Hazard ratio exposure
#> 6 model_1 te Total effect Hazard ratio exposure
#> 7 model_1 pm Proportion mediated Proportion exposure
#> Mediator Outcome Exposure contrast Mediator reference N Case N Non-event N
#> 1 mediator outcome -1.000 -> 1.000 0.002 200 12 188
#> 2 mediator outcome -1.000 -> 1.000 <NA> 200 12 188
#> 3 mediator outcome -1.000 -> 1.000 <NA> 200 12 188
#> 4 mediator outcome -1.000 -> 1.000 <NA> 200 12 188
#> 5 mediator outcome -1.000 -> 1.000 <NA> 200 12 188
#> 6 mediator outcome -1.000 -> 1.000 <NA> 200 12 188
#> 7 mediator outcome -1.000 -> 1.000 <NA> 200 12 188
#> Total follow-up Estimate 95% CI P value Mediator model Outcome model
#> 1 2055.604 0.973 0.333, 2.841 0.960 linear survCox
#> 2 2055.604 0.975 0.332, 2.867 0.964 linear survCox
#> 3 2055.604 0.995 0.941, 1.052 0.854 linear survCox
#> 4 2055.604 0.967 0.333, 2.806 0.951 linear survCox
#> 5 2055.604 1.003 0.960, 1.049 0.880 linear survCox
#> 6 2055.604 0.970 0.333, 2.828 0.956 linear survCox
#> 7 2055.604 0.172 -6.494, 6.838 0.960 linear survCox
if (requireNamespace("regmedint", quietly = TRUE)) {
set.seed(123)
n <- 200
age <- rnorm(n, 60, 8)
exposure_num <- rbinom(n, 1, 0.5)
mediator_num <- rbinom(n, 1, plogis(-0.4 + 0.9 * exposure_num + 0.02 * (age - 60)))
time_event <- rexp(n, rate = exp(-5.8 + 0.55 * exposure_num + 0.45 * mediator_num + 0.02 * (age - 60)))
time_censor <- rexp(n, rate = 0.08)
med_df <- data.frame(
outcome = as.integer(time_event <= time_censor),
outcome_censor = pmax(pmin(time_event, time_censor), 0.1),
exposure = factor(exposure_num, levels = c(0, 1), labels = c("Low risk", "High risk")),
mediator = factor(mediator_num, levels = c(0, 1), labels = c("Low inflammation", "High inflammation")),
age = age
)
res_med <- leo_cox_mediation(
df = med_df, y_out = c("outcome", "outcome_censor"),
x_exp = "exposure", x_med = "mediator", x_cov = "age",
verbose = FALSE
)
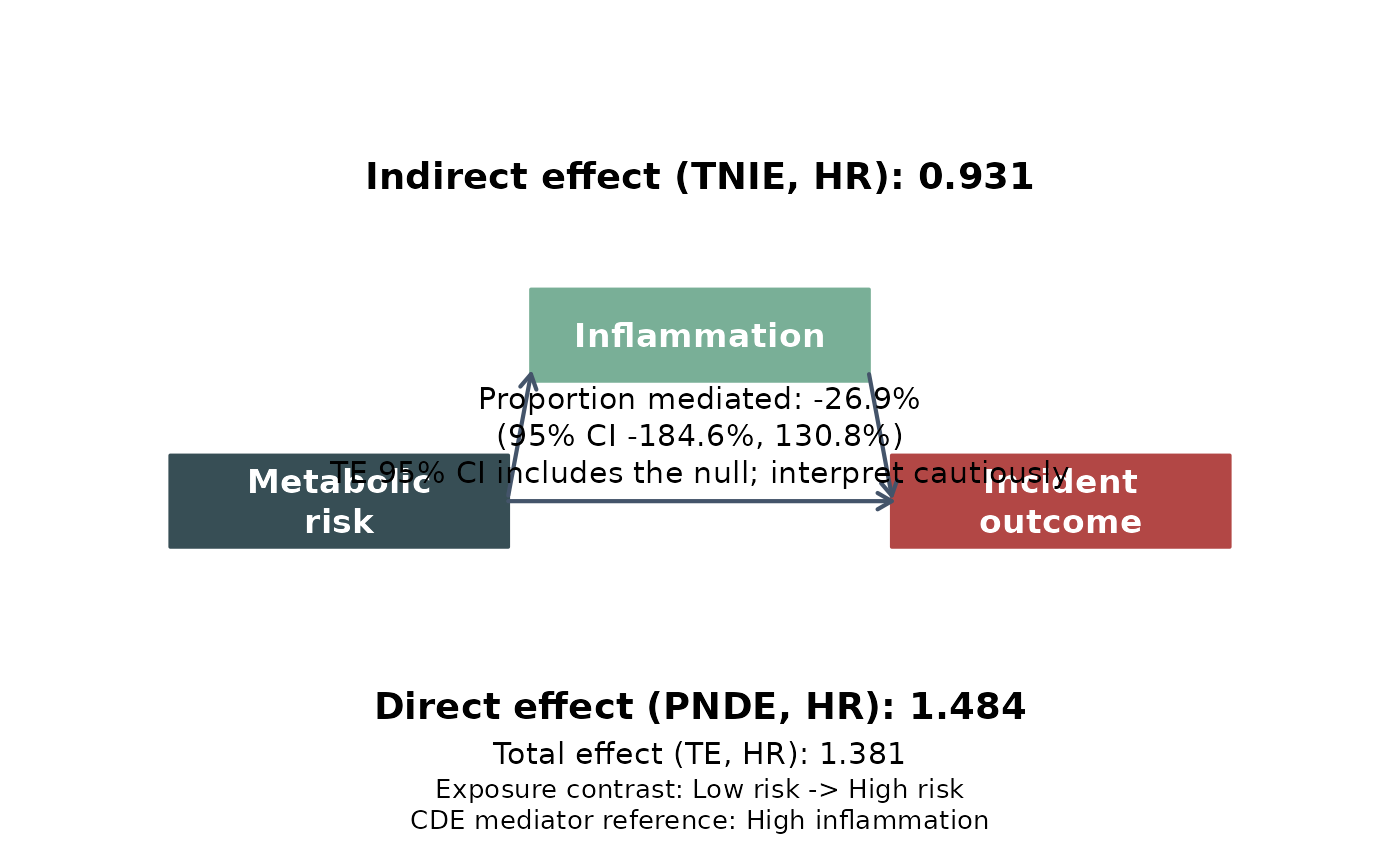
# with default labels and note
leo_cox_mediation_plot(
res_med, model = "model_1",
exposure_label = "Metabolic\nrisk",
mediator_label = "Inflammation",
outcome_label = "Incident\noutcome"
)
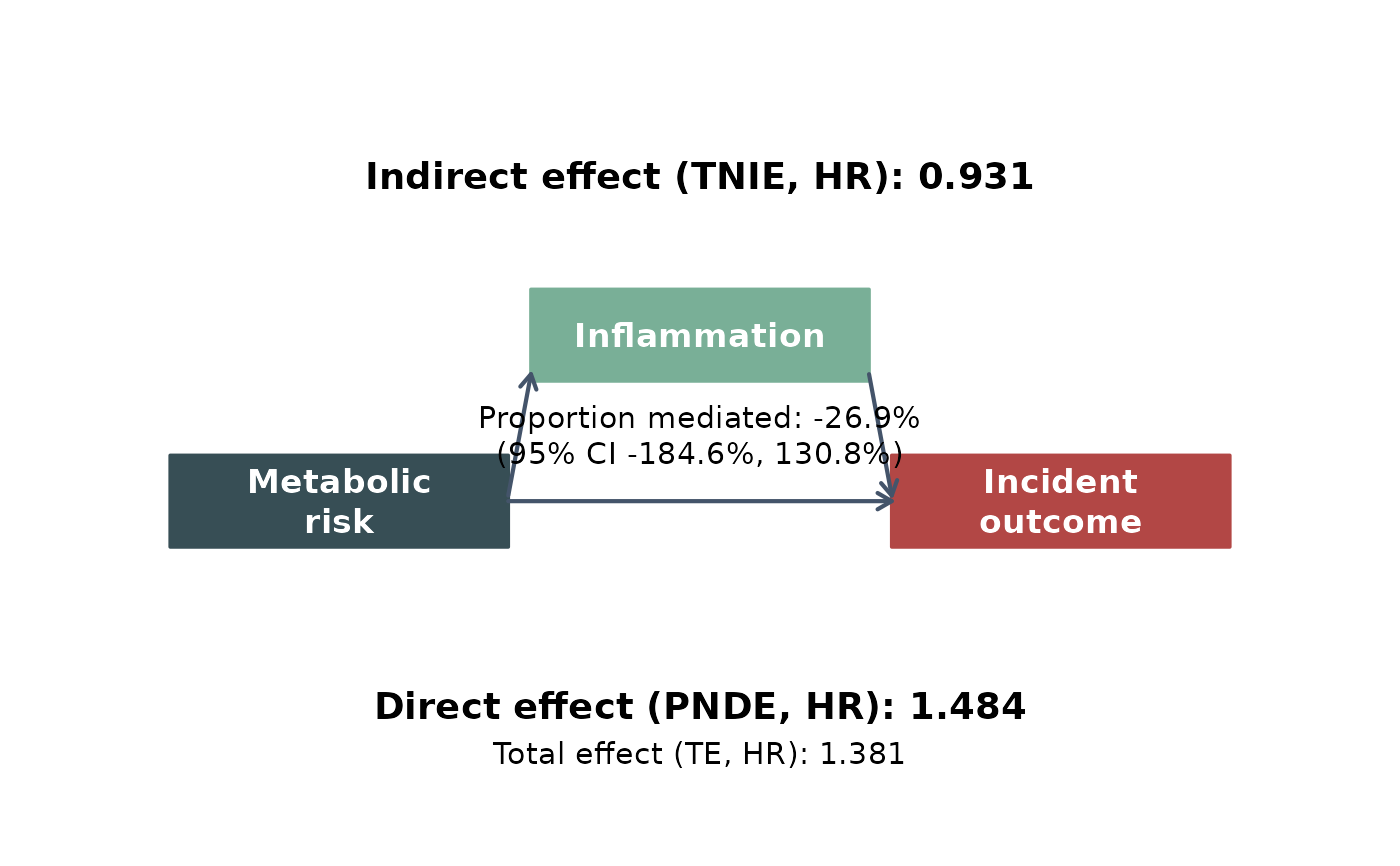
# with no note
leo_cox_mediation_plot(
res_med, model = "model_1",
exposure_label = "Metabolic\nrisk",
mediator_label = "Inflammation",
outcome_label = "Incident\noutcome",
add_note = FALSE
)
}