
Survival Analysis Step 3.1: Mediation Case Study
survival-analysis-step3-1.RmdWhy this article exists
Step 3.0 explains what mediation analysis is for. Step 3.1 is where we turn that into a practical workflow.
The goal here is not just to run leo_cox_mediation(),
but to answer five questions in a way that stays tied to real
output:
- what biological question are we actually asking?
- how do
x_exp_a0,x_exp_a1,x_med_cde, andx_cov_condmap to that question? - when should
mreg = "linear"be preferred overmreg = "logistic"? - what changes when
yreg = "auto"switches fromsurvCoxtosurvAFT_weibull? - how should the scientific interpretation change across those settings?
Build one reproducible scientific story, then vary the scenario
We use a synthetic cohort with the same broad question throughout:
is the association between higher metabolic risk and an incident outcome partly explained by inflammation?
The scenarios differ only in how we encode the mediator and how common the outcome is.
set.seed(20260324)
make_mediation_case <- function(n = 1500, base_log_hazard = -5.8, censor_rate = 0.06) {
age <- rnorm(n, 60, 8)
sex <- factor(sample(c("Female", "Male"), n, replace = TRUE))
smoking <- factor(sample(c("Never", "Ever"), n, replace = TRUE, prob = c(0.6, 0.4)))
sex_num <- as.integer(sex == "Male")
smoking_num <- as.integer(smoking == "Ever")
metabolic_risk_num <- rbinom(
n, 1,
plogis(-0.3 + 0.03 * (age - 60) + 0.30 * sex_num + 0.35 * smoking_num)
)
inflammation_score <- rnorm(
n,
mean = 0.85 * metabolic_risk_num + 0.04 * (age - 60) + 0.35 * smoking_num,
sd = 0.9
)
hazard_rate <- exp(
base_log_hazard +
0.55 * metabolic_risk_num +
0.24 * inflammation_score +
0.02 * (age - 60) +
0.20 * smoking_num
)
event_time <- rexp(n, rate = hazard_rate)
censor_time <- rexp(n, rate = censor_rate)
data.frame(
incident_event = as.integer(event_time <= censor_time),
followup_years = pmin(event_time, censor_time),
metabolic_risk = factor(
metabolic_risk_num,
levels = c(0, 1),
labels = c("Lower risk", "Higher risk")
),
inflammation_score = inflammation_score,
inflammation_group = factor(
as.integer(inflammation_score > 0.5),
levels = c(0, 1),
labels = c("Lower inflammation", "Higher inflammation")
),
age = age,
sex = sex,
smoking = smoking
)
}
rare_case <- make_mediation_case(base_log_hazard = -5.8, censor_rate = 0.06)
common_case <- make_mediation_case(base_log_hazard = -4.6, censor_rate = 0.06)
data.frame(
Scenario = c("Rare-outcome case", "Common-outcome case"),
Participants = c(nrow(rare_case), nrow(common_case)),
Events = c(sum(rare_case$incident_event), sum(common_case$incident_event)),
`Event rate` = c(fmt_pct(mean(rare_case$incident_event)), fmt_pct(mean(common_case$incident_event)))
)
#> Scenario Participants Events Event.rate
#> 1 Rare-outcome case 1500 131 8.7%
#> 2 Common-outcome case 1500 339 22.6%The rare-outcome case will let yreg = "auto" stay on
survCox, because the post-filter, complete-case analysis
set still has an event rate at or below 10%. The common-outcome case is
deliberately built to push the same scientific question onto the AFT
path.
Case 1. A graded biomarker mediator in a rare-outcome setting
This is the most natural scenario when inflammation is scientifically treated as a continuum rather than as a yes/no threshold.
x_med_cde_case1 <- stats::median(rare_case$inflammation_score, na.rm = TRUE)
res_case1 <- leo_cox_mediation(
df = rare_case,
y_out = c("incident_event", "followup_years"),
x_exp = "metabolic_risk",
x_med = "inflammation_score",
x_cov = c("age", "sex", "smoking"),
x_exp_a0 = 0,
x_exp_a1 = 1,
x_med_cde = x_med_cde_case1,
x_cov_cond = list(age = 60, sex = "Female", smoking = "Never"),
mreg = "linear",
yreg = "auto",
interaction = TRUE,
verbose = FALSE
)
case1_eval <- res_case1$evaluation
case1_te <- get_effect_row(res_case1, "te")
case1_tnie <- get_effect_row(res_case1, "tnie")
case1_pm <- get_effect_row(res_case1, "pm")
case1_te_ci <- case1_te[["95% CI"]][1]
case1_tnie_ci <- case1_tnie[["95% CI"]][1]
case1_pm_ci <- case1_pm[["95% CI"]][1]Read the evaluation settings before reading the effects
res_case1$evaluation[, c(
"exposure_contrast", "x_exp_a0", "x_exp_a1",
"mediator_reference", "x_med_cde", "x_cov_cond",
"mreg", "yreg", "interaction"
)]
#> exposure_contrast x_exp_a0 x_exp_a1 mediator_reference x_med_cde
#> 1 Lower risk -> Higher risk 0 1 0.568 0.5680312
#> x_cov_cond mreg yreg interaction
#> 1 age=60.000; sex=Female; smoking=Never linear survCox TRUEThis one table already answers the three user-facing evaluation questions:
-
x_exp_a0 = 0,x_exp_a1 = 1means we are comparingLower risk -> Higher risk -
x_med_cde = 0.568means the controlled direct effect is being evaluated at the median inflammation score -
x_cov_cond = age=60; sex=Female; smoking=Nevermeans the reported effects are evaluated for that covariate profile, not as a profile-free population average
In plain language, the CDE here asks:
if two otherwise similar 60-year-old never-smoking women differed only in metabolic-risk status, but both were fixed at an inflammation score of 0.568, how different would their outcome risk be?
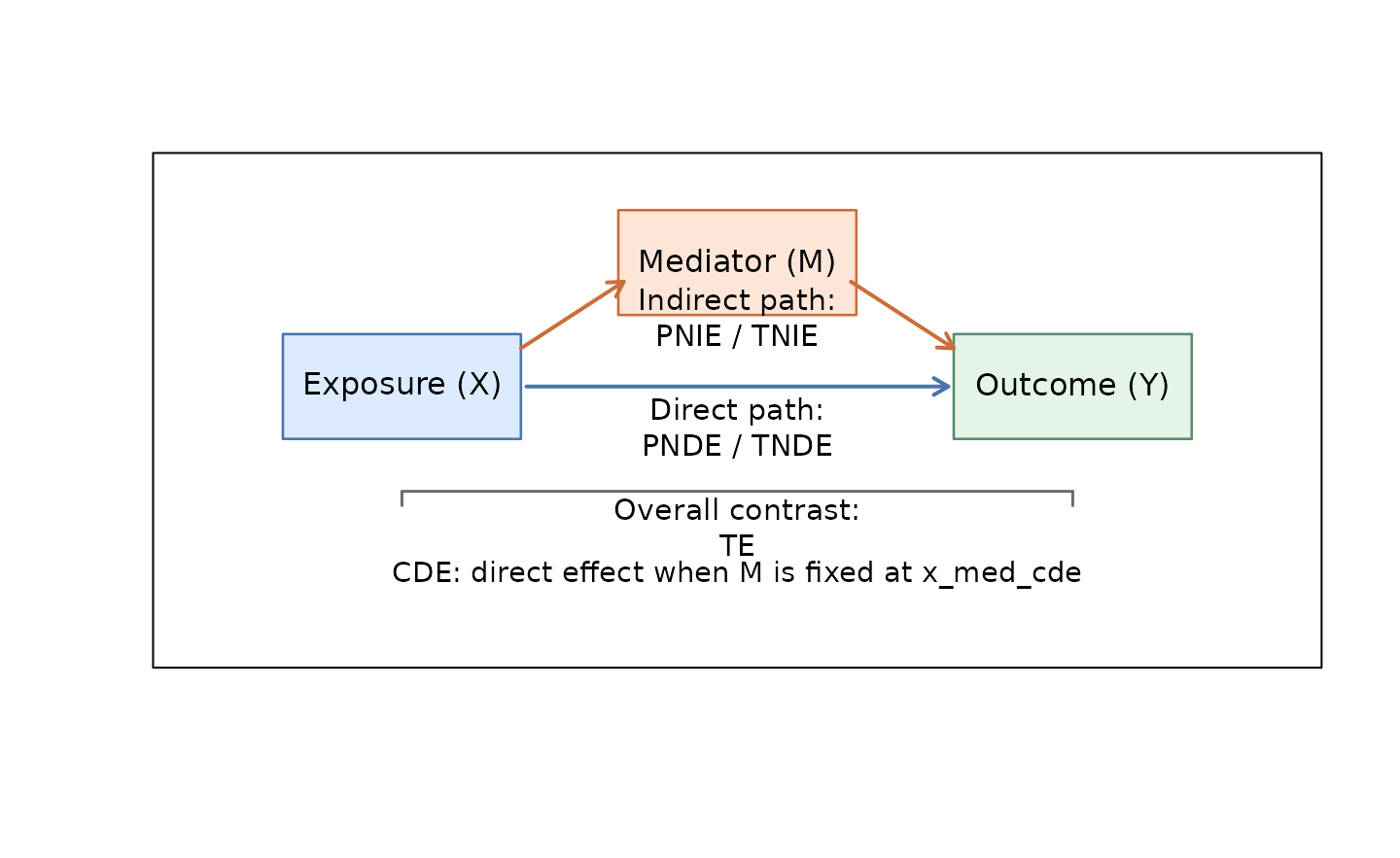
Keep one simple map of the effect names in mind
| Code | Full name | How to read it |
|---|---|---|
| TE | Total Effect | the overall association |
| TNIE | Total Natural Indirect Effect | the part that travels through the mediator |
| PNDE | Pure Natural Direct Effect | the part that does not travel through the mediator |
| PM | Proportion Mediated | the share of the total effect attributed to the mediator pathway |
| CDE | Controlled Direct Effect | the direct effect when the mediator is fixed at x_med_cde |
| TNDE | Total Natural Direct Effect | an alternative natural direct-effect definition |
| PNIE | Pure Natural Indirect Effect | an alternative natural indirect-effect definition |
For most papers, the first pass is usually TE,
TNIE, PNDE, and PM.
CDE becomes important when the scientific question is
explicitly about fixing the mediator at a chosen level.
That is also why Mediator reference appears only in the
cde row: only the controlled direct effect needs a fixed
mediator value.
Visualize the biology before over-interpreting the decomposition
First look at whether the mediator really behaves like a graded biomarker across the two exposure groups.
ggplot2::ggplot(
rare_case,
ggplot2::aes(x = metabolic_risk, y = inflammation_score, fill = metabolic_risk)
) +
ggplot2::geom_violin(trim = FALSE, alpha = 0.35, color = NA) +
ggplot2::geom_boxplot(width = 0.16, outlier.alpha = 0.15, color = "#1f1f1f") +
ggplot2::scale_fill_manual(values = c("#5B8FF9", "#D14A61")) +
ggplot2::labs(
title = "Inflammation score is shifted upward in the higher-risk group",
x = "Metabolic risk group",
y = "Inflammation score"
) +
ggplot2::theme_minimal(base_size = 12) +
ggplot2::theme(legend.position = "none")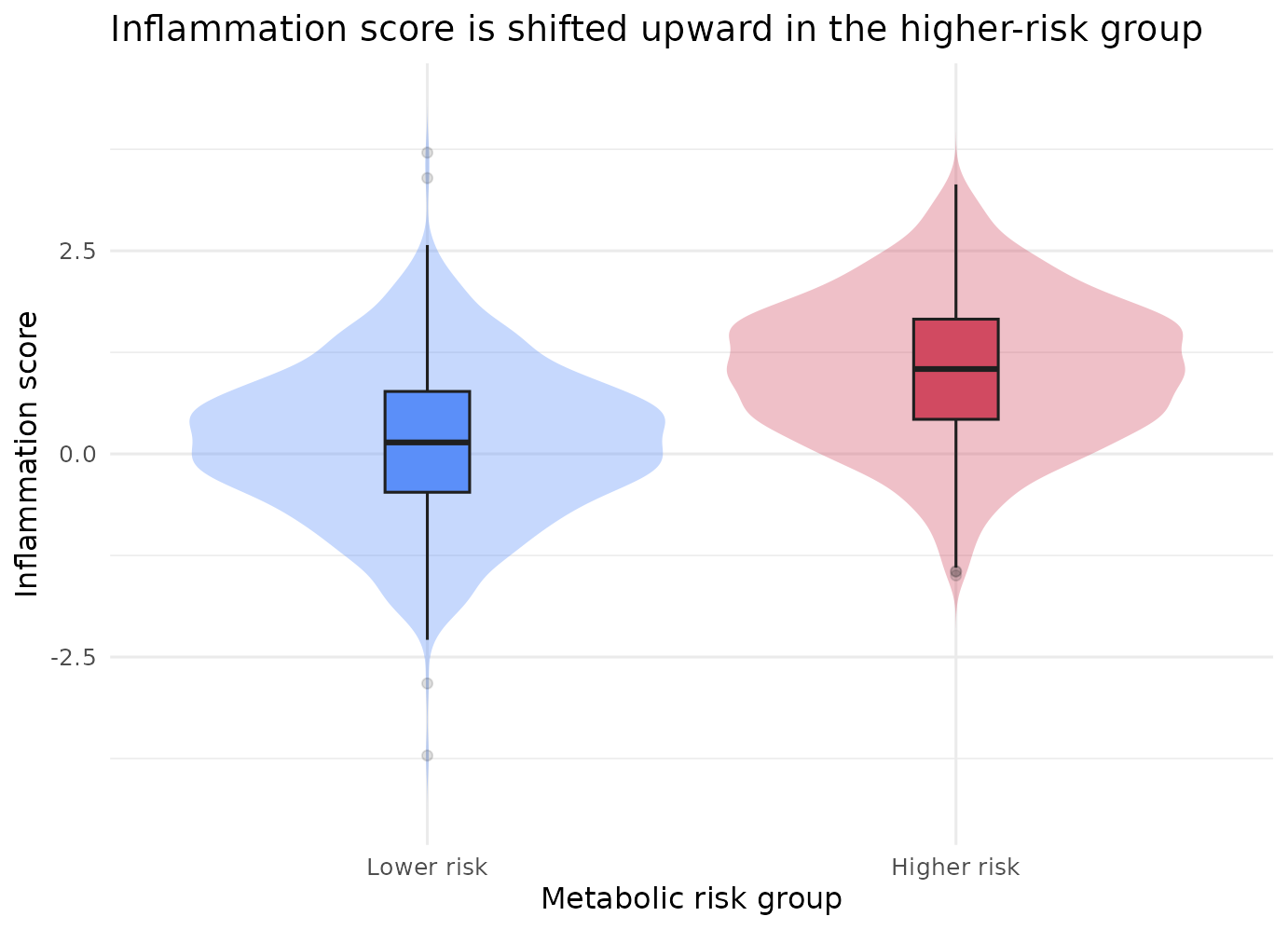
Now look at the crude joint pattern that motivates keeping
interaction = TRUE in the outcome model.
joint_event_case1 <- dplyr::summarise(
dplyr::group_by(rare_case, metabolic_risk, inflammation_group),
n = dplyr::n(),
event_rate = mean(incident_event),
.groups = "drop"
)
ggplot2::ggplot(
joint_event_case1,
ggplot2::aes(x = inflammation_group, y = event_rate, fill = metabolic_risk)
) +
ggplot2::geom_col(position = ggplot2::position_dodge(width = 0.7), width = 0.62) +
ggplot2::geom_text(
ggplot2::aes(label = paste0(fmt_pct(event_rate), "\n(n=", n, ")")),
position = ggplot2::position_dodge(width = 0.7),
vjust = -0.15,
size = 3.3
) +
ggplot2::scale_fill_manual(values = c("#5B8FF9", "#D14A61")) +
ggplot2::scale_y_continuous(
labels = function(x) paste0(round(100 * x), "%"),
limits = c(0, max(joint_event_case1$event_rate) * 1.18)
) +
ggplot2::labs(
title = "The highest crude event rate appears in the higher-risk plus higher-inflammation stratum",
x = "Inflammation group",
y = "Observed event rate",
fill = "Metabolic risk"
) +
ggplot2::theme_minimal(base_size = 12)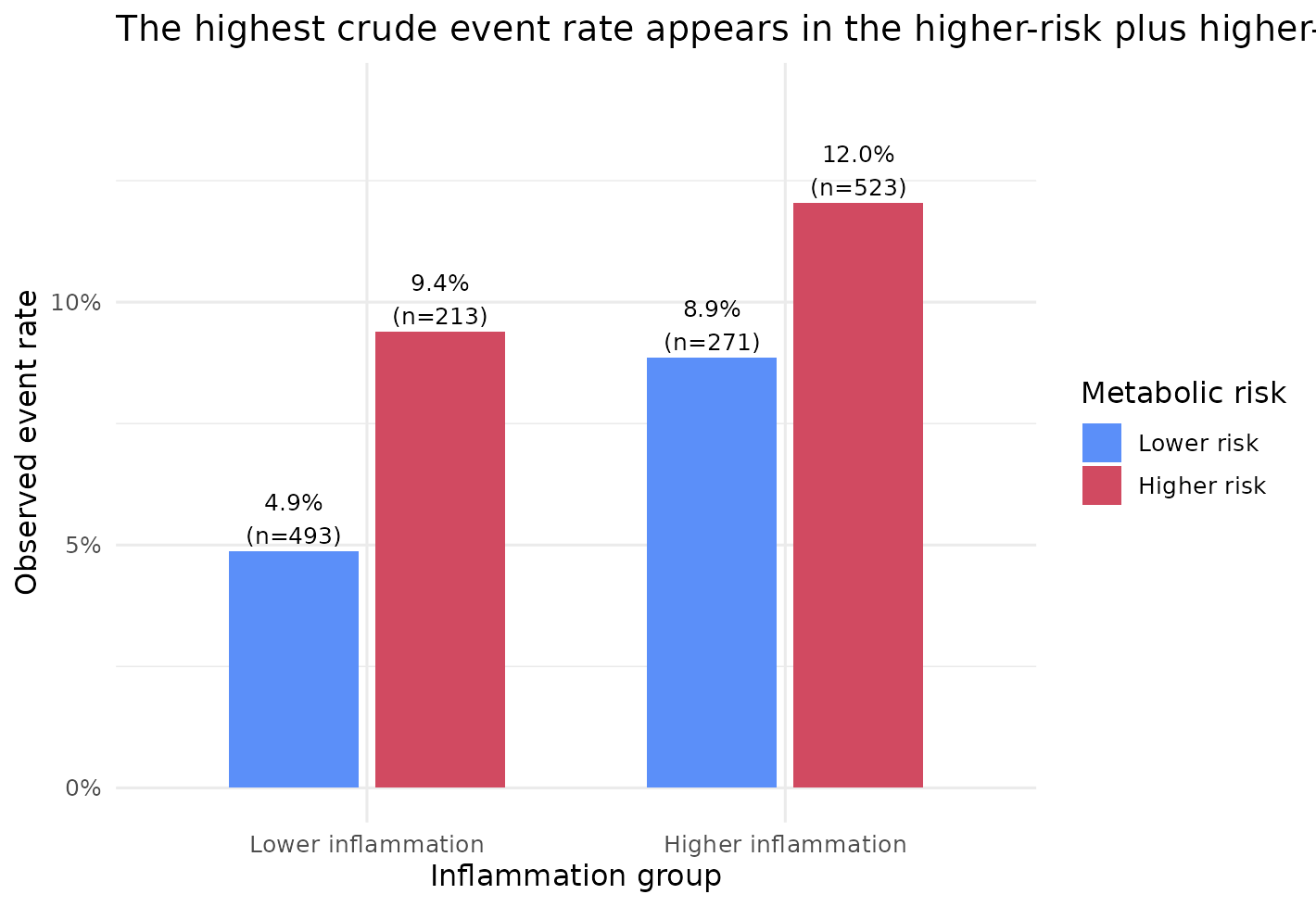
That figure does not prove interaction by itself, but it shows why
the default interaction = TRUE is often a sensible first
pass: it avoids forcing the effect of metabolic risk to be identical at
every inflammation level.
Interpret the actual mediation result, not an imagined one
dplyr::filter(res_case1$result, `Effect code` %in% c("te", "tnie", "pm"))
#> Model Effect code Effect Scale Exposure
#> 1 model_1 tnie Total natural indirect effect Hazard ratio metabolic_risk
#> 2 model_1 te Total effect Hazard ratio metabolic_risk
#> 3 model_1 pm Proportion mediated Proportion metabolic_risk
#> Mediator Outcome Exposure contrast
#> 1 inflammation_score incident_event Lower risk -> Higher risk
#> 2 inflammation_score incident_event Lower risk -> Higher risk
#> 3 inflammation_score incident_event Lower risk -> Higher risk
#> Mediator reference N Case N Non-event N Total follow-up Estimate
#> 1 <NA> 1500 131 1369 23086.95 1.246
#> 2 <NA> 1500 131 1369 23086.95 1.877
#> 3 <NA> 1500 131 1369 23086.95 0.423
#> 95% CI P value Mediator model Outcome model
#> 1 1.018, 1.525 0.033 linear survCox
#> 2 1.303, 2.703 <0.001 linear survCox
#> 3 0.027, 0.820 0.036 linear survCoxBecause this rare-outcome scenario stayed on survCox,
the ratio scale is a hazard ratio.
In this fitted example:
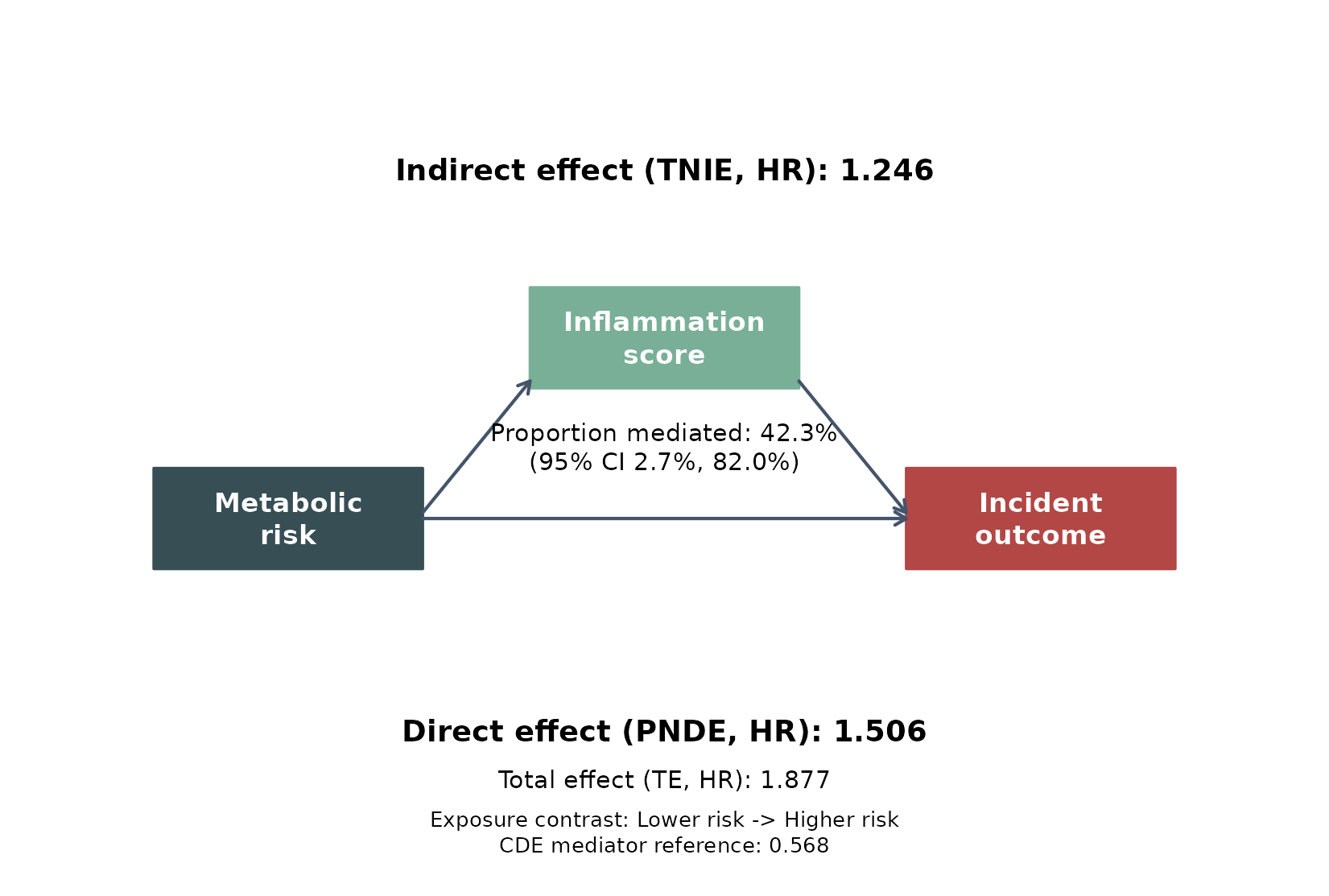
- the total effect was 1.877 (1.303, 2.703), meaning the higher-risk group had a substantially higher incident hazard overall
- the indirect pathway through the continuous inflammation score was 1.246 (1.018, 1.525), so the mediator pathway itself still points in the same harmful direction
- the estimated proportion mediated was 42.3% (0.027, 0.820), suggesting that inflammation may explain a sizeable share of the total association in this synthetic cohort
Here the pm confidence interval stays above
0, so this particular simulated example is more consistent
with a positive mediated share than with a null mediated
contribution.
Scientifically, that is the pattern we would describe as:
higher metabolic risk is associated with both a higher inflammation burden and a higher incident hazard, and the mediation decomposition is compatible with inflammation carrying a meaningful part of that pathway.
The pathway figure makes the same result easier to communicate in a paper or slide deck.
leo_cox_mediation_plot(
x = res_case1,
exposure_label = "Metabolic\nrisk",
mediator_label = "Inflammation\nscore",
outcome_label = "Incident\noutcome",
language = "en",
palette = "jama"
)
Case 2. The same biology, but a thresholded binary mediator
Sometimes the scientific question is about a thresholded inflammatory
state rather than a graded biomarker. The model then changes from
mreg = "linear" to mreg = "logistic".
res_case2 <- leo_cox_mediation(
df = rare_case,
y_out = c("incident_event", "followup_years"),
x_exp = "metabolic_risk",
x_med = "inflammation_group",
x_cov = c("age", "sex", "smoking"),
x_exp_a0 = 0,
x_exp_a1 = 1,
x_med_cde = "Higher inflammation",
x_cov_cond = list(age = 60, sex = "Female", smoking = "Never"),
mreg = "logistic",
yreg = "auto",
interaction = TRUE,
verbose = FALSE
)
case2_tnie <- get_effect_row(res_case2, "tnie")
case2_pm <- get_effect_row(res_case2, "pm")
dplyr::bind_rows(
case_summary_row(res_case1, "Case 1: rare outcome + continuous mediator"),
case_summary_row(res_case2, "Case 2: rare outcome + binary mediator")
)
#> Scenario Event rate Outcome model
#> 1 Case 1: rare outcome + continuous mediator 8.7% survCox
#> 2 Case 2: rare outcome + binary mediator 8.7% survCox
#> Total effect Indirect effect Proportion mediated
#> 1 1.877 (1.303, 2.703) 1.246 (1.018, 1.525) 42.3% (2.7%, 82.0%)
#> 2 1.878 (1.304, 2.706) 1.100 (0.920, 1.315) 19.5% (-16.4%, 55.4%)The scientific lesson is not that binary mediators are wrong. It is
that the mediator encoding should match the question. For logistic
mediators, x_med_cde should now be given as the original
observed mediator level, not an internal 0/1 recode:
- in Case 1, the indirect pathway was 1.246 on the hazard-ratio scale,
with
pmestimated at 42.3% - in Case 2, the same broad story became much weaker and less precise
(
tnie= 1.100,pm= 19.5%)
That is a useful teaching contrast. If the biology is genuinely graded, turning the mediator into a yes/no label can discard pathway information. If the real question is threshold-based, however, the binary mediator may be the better scientific choice even if it is statistically less efficient.
Case 3. The same pathway question, but with a more common outcome
Now keep the same exposure-mediator story and only make the event
more common. This is where yreg = "auto" becomes
important.
res_case3 <- leo_cox_mediation(
df = common_case,
y_out = c("incident_event", "followup_years"),
x_exp = "metabolic_risk",
x_med = "inflammation_score",
x_cov = c("age", "sex", "smoking"),
x_exp_a0 = 0,
x_exp_a1 = 1,
x_med_cde = stats::median(common_case$inflammation_score, na.rm = TRUE),
x_cov_cond = list(age = 60, sex = "Female", smoking = "Never"),
mreg = "linear",
yreg = "auto",
interaction = TRUE,
verbose = FALSE
)
case3_te <- get_effect_row(res_case3, "te")
case3_tnie <- get_effect_row(res_case3, "tnie")
case3_pm <- get_effect_row(res_case3, "pm")
res_case3$evaluation[, c("event_rate", "mreg", "yreg", "interaction", "x_cov_cond")]
#> event_rate mreg yreg interaction
#> 1 0.226 linear survAFT_weibull TRUE
#> x_cov_cond
#> 1 age=60.000; sex=Female; smoking=Never
dplyr::filter(res_case3$result, `Effect code` %in% c("te", "tnie", "pm"))
#> Model Effect code Effect Scale Exposure
#> 1 model_1 tnie Total natural indirect effect Time ratio metabolic_risk
#> 2 model_1 te Total effect Time ratio metabolic_risk
#> 3 model_1 pm Proportion mediated Proportion metabolic_risk
#> Mediator Outcome Exposure contrast
#> 1 inflammation_score incident_event Lower risk -> Higher risk
#> 2 inflammation_score incident_event Lower risk -> Higher risk
#> 3 inflammation_score incident_event Lower risk -> Higher risk
#> Mediator reference N Case N Non-event N Total follow-up Estimate
#> 1 <NA> 1500 339 1161 19723.5 0.860
#> 2 <NA> 1500 339 1161 19723.5 0.396
#> 3 <NA> 1500 339 1161 19723.5 0.107
#> 95% CI P value Mediator model Outcome model
#> 1 0.754, 0.981 0.024 linear survAFT_weibull
#> 2 0.303, 0.518 <0.001 linear survAFT_weibull
#> 3 -0.011, 0.225 0.076 linear survAFT_weibullHere the event rate was 22.6%, so yreg = "auto" switched
to survAFT_weibull.
That changes the interpretation scale:
-
te= 0.396 is now a time ratio, not a hazard ratio - because the time ratio is below 1, the higher-risk group has shorter event-free time
-
tnie= 0.860 means the inflammation pathway is also pointing toward shorter event-free time
This is the same biological story on a different model scale. The key
practical point is that the wording must change with
yreg:
-
survCox: talk about higher or lower hazard -
survAFT_weibull: talk about shorter or longer event-free time
Put the three scenarios side by side
dplyr::bind_rows(
case_summary_row(res_case1, "Case 1: rare outcome + continuous mediator"),
case_summary_row(res_case2, "Case 2: rare outcome + binary mediator"),
case_summary_row(res_case3, "Case 3: common outcome + continuous mediator")
)
#> Scenario Event rate Outcome model
#> 1 Case 1: rare outcome + continuous mediator 8.7% survCox
#> 2 Case 2: rare outcome + binary mediator 8.7% survCox
#> 3 Case 3: common outcome + continuous mediator 22.6% survAFT_weibull
#> Total effect Indirect effect Proportion mediated
#> 1 1.877 (1.303, 2.703) 1.246 (1.018, 1.525) 42.3% (2.7%, 82.0%)
#> 2 1.878 (1.304, 2.706) 1.100 (0.920, 1.315) 19.5% (-16.4%, 55.4%)
#> 3 0.396 (0.303, 0.518) 0.860 (0.754, 0.981) 10.7% (-1.1%, 22.5%)This comparison is useful because it separates three different decisions:
- how to define the mediator scientifically
- how much information is lost when a graded mediator is dichotomized
- how the interpretation scale changes when the outcome is no longer rare
Key practical takeaways
If you only keep six practical rules from Step 3.1, make them these:
- choose
x_exp_a0andx_exp_a1to match the exposure contrast you would actually write in a Results section - choose
x_med_cdeto represent the mediator level at which you want the controlled direct effect to be interpreted; for continuous biomarkers, the median is often a clean first choice - choose
x_cov_condto represent a real covariate profile, especially whenx_covincludes factor variables - use
mreg = "linear"when the mediator is scientifically graded, andmreg = "logistic"when the mediator question is genuinely threshold-based - let
yreg = "auto"protect you from usingsurvCoxin obviously non-rare settings, but always read the finalOutcome modelbefore interpreting the ratio scale - write the scientific conclusion from the actual fitted output, not from the existence of a pathway diagram or a single positive point estimate